
Biology has changed dramatically over the past two decades, making engineering based on biological systems possible. The genomic revolution that gave us the ability to sequence the genetic code (DNA) of our cells was a major driver of this dramatic change. One of the latest discoveries brought about by the genomic revolution is the ability to use CRISPR to precisely edit DNA in the body.
High-level expressions of the genetic code, such as protein synthesis, are called phenotypes. High-throughput phenotypic data combined with precise editing of DNA connects changes in the underlying code to external phenotypes.
◆The potential of synthetic biology
Synthetic biology will have a transformative impact on food, energy, climate, medicine and materials… and every field in the world.
Synthetic biology has already given the world the ability to obtain pig insulin without sacrificing the pig (something that was possible in previous stages of genetic engineering), synthetic leather, coats that are not made of spider silk at all, and anti-malarial and anti-cancer Medicines, meatless burgers that taste like meat, renewable biofuels, hoppy beers without the hops, the scent of extinct flowers, artificial collagen for cosmetics, eliminating the genes of dengue-carrying mosquitoes. Many believe this is just the tip of the iceberg, as the ability to engineer organisms brings endless possibilities to transform the world, and the level of public and private investment in this area continues to grow.
In addition, after entering the third wave of AI, AI focuses on integrating the environment into the model, and its potential to impact synthetic biology has greatly increased.
It is well known that the genotype of an organism is not so much a blueprint for its phenotype but rather the initial conditions of a complex, interconnected, dynamic system. Biologists have spent decades constructing and managing a large set of properties—regulations, correlations, rates of change, and function—that describe this complex, dynamic system. Other resources such as gene networks, known functional associations, protein-protein interactions, protein-metabolite interactions, and knowledge-driven dynamic models of transcription, translation, and interactions provide rich resources for artificial intelligence models.
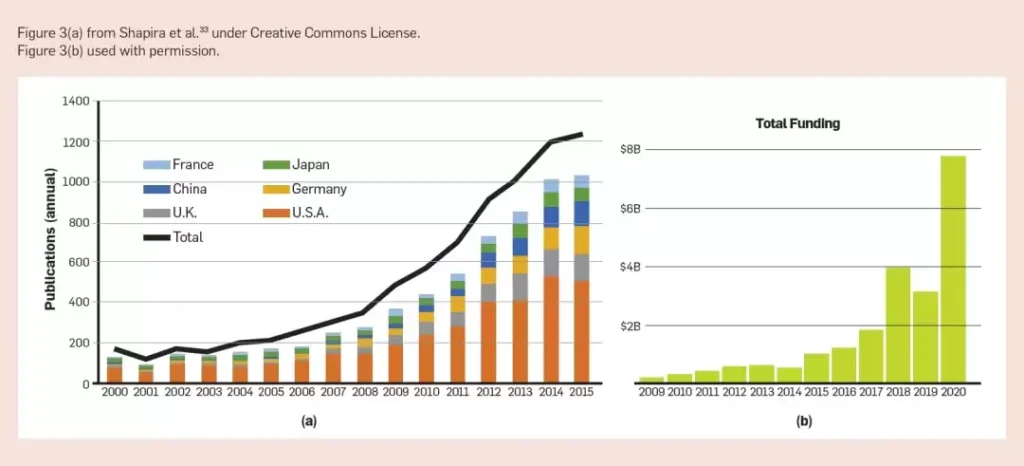
Model interpretability is also critical for uncovering new design principles. These models give biologists the ability to solve more complex questions about biological systems and build comprehensive, interpretable models to accelerate discovery and research. The growth in knowledge and resources in the field is evident in the number of synthetic biology publications and the commercial opportunities in synthetic biology.
◆AI and its impact on synthetic biology
Compared with the potential of AI in the field of synthetic biology, its impact in the field of synthetic biology is limited.
We have seen successful applications of AI, but it is still limited to specific data sets and research questions. The current challenge for AI in the field remains how general it is to a wider range of applications and other data sets.
Data mining, statistics, and mechanistic modeling are currently the main drivers of computational biology and bioinformatics in the field, but the boundaries between these techniques and artificial intelligence/machine learning are often blurred. For example, clustering is a data mining technique that can identify patterns and structures in gene expression data that can indicate whether engineering modifications lead to toxic outcomes in cells. These clustering techniques can also serve as unsupervised learning models to find structure in unlabeled data sets. These classic technologies and new AI/ML (machine learning) methods under development will play a greater role and impact in the field of synthetic biology in the future, because by then people will become accustomed to larger data sets. Transcriptome data volume is doubling every 7 months, and high-throughput workflows for proteomics and metabolomics are increasingly available.
In addition, the progressive automation and miniaturization of microfluidic chips for laboratory work indicates that data processing and analysis will double the productivity of synthetic biology in the future. DARPA’s Collaborative Discovery and Design (SD2, 2018–2021) program focuses on building artificial intelligence models and aims to bridge the gap between AI and synthetic biology needs. This is also evident in some companies adopting SoTA technology in this field (such as Amyris, Zymergen or Ginkgo Bioworks).
AI and synthetic biology overlap in some ways, such as applying existing AI/ML to existing datasets; generating new datasets (such as the upcoming NIH Bridge2AI); and creating new AI/ML technologies to apply New or existing data. Although SD2 has contributed to the last item, it still has certain potential and has a long way to go in the future.
Artificial intelligence can help synthetic biology overcome a big challenge: predicting the impact of bioengineering methods on biological subjects and the environment. Since the results of bioengineering cannot be predicted, the cell engineering goals of synthetic biology (i.e., inverse design) can only be achieved through a lot of trial and error. Artificial intelligence provides an opportunity to use publicly available and experimental data to predict impacts on biological agents and the environment.
Designing genetic structures for cell programming. Much research in the field of synthetic biology focuses on the engineering of genetic structures/gene circuits, which is very different from the challenges faced in designing electronic circuits.
Artificial intelligence technology, which combines known biophysical, machine learning and reinforcement learning models, can effectively predict the impact of structures on subjects and vice versa. Although it is already quite powerful, there is still room for improvement. In terms of machine-assisted gene circuit design, various artificial intelligence technologies have been put into application, including expert systems, multi-agent systems, constrained reasoning, heuristic search, optimization and machine learning.
Sequence-based models and graph convolutional networks have also received attention in the field of engineered biological systems. Factor-graph neural networks have been used to incorporate biological knowledge into deep learning models. Graph convolutional networks have been used to predict protein function from protein-protein interaction networks. Sequence-based convolutional and recurrent neural network models have been used to identify potential binding sites for proteins, gene expression, and the design of new biological structures. Artificial intelligence is most useful when applied to the development of comprehensive models, which will reduce the number of experiments or designs that need to be performed.
Metabolic engineering. In metabolic engineering, artificial intelligence has been applied to almost all stages of the bioengineering process. For example, artificial neural networks have been used to predict translation start sites, annotate protein functions, predict synthetic pathways, and optimize the expression levels of multiple foreign genes. , predict the strength of regulatory elements, predict plasmid expression, optimize nutrient concentration and fermentation conditions, predict enzyme kinetic parameters, understand the correlation between genotype and phenotype, predict the guidance effect of CRISPR, etc. Clustering has been used to discover secondary metabolite biosynthetic gene clusters and identify enzymes that catalyze specific reactions. Ensemble methods have been used to predict pathway dynamics, optimal growth temperatures, and to find proteins conferring higher fitness in directed evolution approaches. Support vector machines have been used to optimize ribosome binding site sequences and predict the behavior of CRISPR guide RNAs. Among the various stages of metabolic engineering, AI is most promising for application in process scale-up, which is a significant bottleneck in the field, and in downstream processing (e.g., systematic extraction of produced molecules from fermentation broth).
Experiment automation. The impact of AI extends well beyond the “learning” phase of the DBTL cycle, in helping automate laboratory work and recommend experimental designs. Automation is becoming increasingly important in practice because it is the most reliable way to obtain the high-quality, high-volume, low-bias data needed to train artificial intelligence algorithms, and it also enables predictable bioengineering. Automation provides the opportunity to quickly transfer and expand complex protocols to other laboratories. For example, liquid handling robotic stations form the backbone of biofoundries and cloud laboratories. These foundries can already see themselves being disrupted by robots and planning algorithms in the future, gaining the ability to iterate quickly through DBTL cycles. Semantic webs, ontologies, and schemas revolutionize the representation, communication, and exchange of designs and protocols. These tools enable rapid experimentation and generate more data in a structured, queryable format. In a field where most content is either lost or manually recorded in lab notes, the promise of artificial intelligence is driving significant changes in the field, reducing barriers to generating data.
Microfluidics is an alternative to macroscopic liquid handling with higher throughput, less reagent consumption, and cheaper fouling. In fact, microfluidics may be a key technology for realizing self-driving laboratories, and it is expected to greatly speed up the research and development process by using artificial intelligence to enhance automated experimental platforms. The autonomous driving laboratory involves a fully automated DBTL cycle, in which artificial intelligence algorithms actively search for promising experimental procedures by making hypotheses based on previous experimental results. So this may be the biggest opportunity for artificial intelligence researchers in synthetic biology. While automated DBTL loops have already been demonstrated in liquid-handling robotic workstations, the scalability, high-throughput capabilities, and manufacturing flexibility offered by microfluidic chips may provide the final technological leap that makes artificial intelligence a reality.
◆Challenges facing using AI to study synthetic biology
Artificial intelligence has begun to make its way into various synthetic biology applications, but technical and social issues remain as barriers between the two fields.
Technical challenges. The technical challenges of applying artificial intelligence to synthetic biology are: data is scattered across different modalities, difficult to combine, unstructured, and often lacks the context in which the data was collected; models require much more data than is typically collected in a single experiment There are many, and there is a lack of interpretability and uncertainty quantification; and there are no metrics or standards to effectively evaluate the performance of the model in the context of larger design tasks. Furthermore, experiments are often designed to explore only positive outcomes, which complicates or biases the evaluation of models.
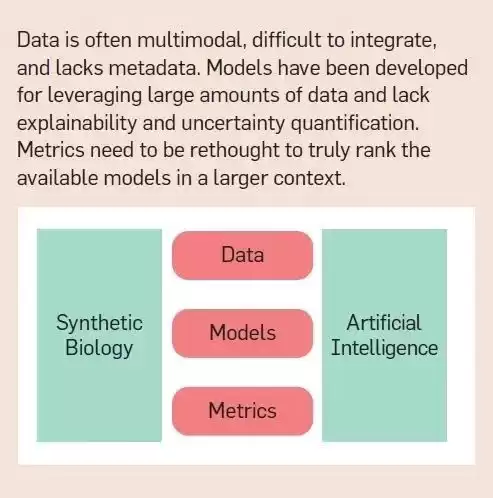
Data challenges. The lack of suitable data sets remains the primary obstacle to integrating artificial intelligence with synthetic biology. Applying artificial intelligence to synthetic biology requires large amounts of labeled, curated, high-quality, context-rich data from individual experiments. Although the community has made progress in building databases containing sequences (even whole genomes) and phenotypes of various organisms, marker data are still scarce. By “marker data” we mean phenotypic data mapped to measurements that capture their biological function or cellular response. It’s the presence of this kind of measurement and labeling that has allowed AI/ML and synthetic biology solutions to mature, as in other fields, pitting AI against human capabilities.
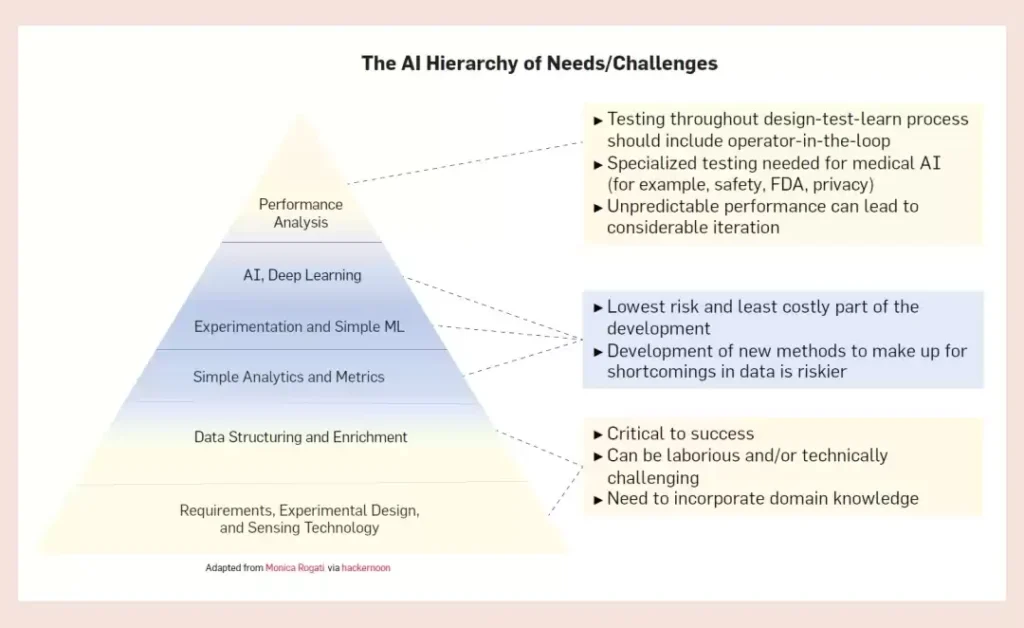
Lack of investment in data engineering is part of the reason for the lack of suitable data sets. Overshadowed by advances in AI technology, the computing infrastructure needs that support and ensure its success are often lost sight of. The AI community calls this the pyramid of needs, and data engineering is an important part of it.
Data engineering includes the steps of experiment planning, data collection, structuring, access and exploration. Successful AI application stories include standardized, consistent, and replicable data engineering steps. While we can now collect biological data at unprecedented scale and detail, this data is often not immediately applicable to machine learning. There are still many barriers to the adoption of community-wide standards for storing and sharing measurement data, experimental conditions, and other metadata that make data more amenable to AI technologies. Rigorous work and a high level of consensus will be needed to enable rapid adoption of these standards while promoting common standards for data quality assessment. In short, AI models require consistent and comparable measurements across all experiments, which can extend experimental timelines. This requirement adds a huge burden to scientific experimenters who already follow complex protocols. As a result, the long-term need to collect data is often sacrificed in order to meet looming project deadlines.
This situation often results in sparse data sets that represent only a small portion of the multiple layers that make up the omics data stack. In this case, data representation has a significant impact on the ability to integrate these isolated data sets for comprehensive modeling. The industry is currently investing heavily in various verticals to perform data cleansing, schema alignment, and extract, transform, and load operations (ETL) to collect unmanageable digital data and prepare it into a form suitable for analysis . These tasks take up nearly 50% to 80% of data scientists’ time, limiting their ability to explore deeply. Handling a large number of data types (data multimodality) is a challenge for synthetic biology researchers, and the complexity of preprocessing activities increases dramatically with the increase in data diversity compared to the data volume.
Modeling/Algorithmic Challenges. Many of the popular algorithms driving current advances in artificial intelligence (such as those in computer vision and NLP) are not robust when analyzing omics data. Traditional applications of these models often suffer from the “curse of dimensionality” when applied to data collected in specific experiments. Under certain conditions, a single experimenter can generate more than 12,000 measurements (dimensions) of genomic, transcriptomic, and proteomic data for an organism. For such an experiment, the number of labeled instances (e.g., success or failure) is usually only a few dozen to a few hundred at most. For these high-dimensional data types, the dynamics of the system (temporal resolution) are rarely captured. These measurement errors make inferences about complex dynamic systems a significant challenge.
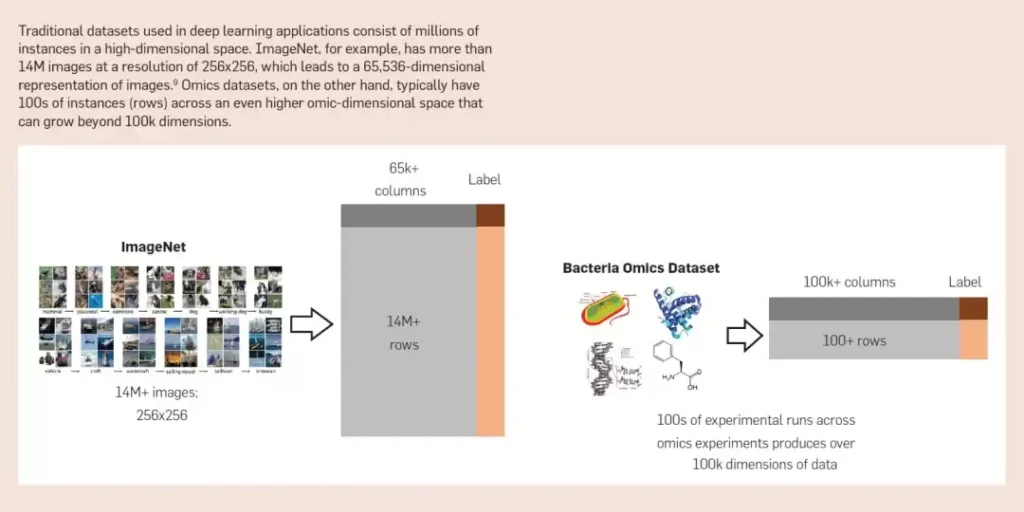
Omics data have both similarities and differences with other data modalities such as sequential, textual, and network-based data, and classical approaches are not always applicable. Common features of these data include positional encodings and dependencies, as well as complex interaction patterns. However, there are some fundamental differences between these data, such as: their underlying representation, the context required for meaningful analysis, and the relevant normalization across modalities for biologically meaningful comparisons. Therefore, it is difficult to find robust generative models (similar to Gaussian models or stochastic block models) that can accurately describe omics data.
Furthermore, biological sequences and systems represent complex biological functional codes, but there are few systematic approaches to interpreting these codes in a manner similar to interpreting semantics or from the context of written texts. These different characteristics make it challenging to extract insights, generate and validate hypotheses through data exploration. Engineering biology involves the challenge of learning black-box systems, where we can observe the inputs and outputs but our understanding of the inner workings of the system is limited. Considering that these biological systems operate in a combinatorial large parameter space, artificial intelligence solutions using strategies to effectively design experiments to explore biological systems to generate various hypotheses and test them equals a huge need in this space and Chance.
Finally, many popular AI algorithm solutions do not explicitly account for uncertainty and do not show robust mechanisms for controlling errors under input perturbations. This fundamental gap is particularly important in the synthetic biological space given the inherent stochasticity and noise in the biological systems we are trying to design.
Metrics/Evaluation Challenges. Standard AI evaluation metrics based on prediction and accuracy are insufficient for application in the field of synthetic biology. Metrics based on the accuracy of regression models like ℝ or classification models cannot account for the complexity of the underlying biological systems we are trying to model. In this field, it is equally important to quantify other indicators of a model’s ability to illuminate the inner workings of biological systems and capture existing domain knowledge. To this end, AI solutions that incorporate principles of explainability and transparency are key to supporting iterative and interdisciplinary research. Furthermore, the ability to appropriately quantify uncertainty requires that we creatively develop new metrics to measure the effectiveness of these methods.
We also need appropriate experimental design metrics. Evaluating and validating models in synthetic biology sometimes requires additional experiments and additional resources. A small number of misclassifications or minor errors can have a significant impact on the research objectives. These costs should be integrated into the objective function or evaluation of the AI model to reflect the real-world impact of misclassification.
Sociological Challenges. Sociological issues may be more challenging than technical barriers to leveraging artificial intelligence combined with synthetic biology (and vice versa). Our impression is that a lack of coordination and understanding between the disparate cultures involved in the study leads to some sociological obstacles. Although there are already some solutions to this obstacle, it is interesting to note that there are still some long-standing sociological problems in academia and industry.
Social problems arise because two very different groups of experts: computational scientists and laboratory scientists, collide and create certain differences in their work.
There are so many differences in the training that computational scientists and laboratory scientists receive. Computational scientists by training tend to focus on abstraction, with a passion for automation, computational efficiency, and disruptive approaches. They naturally tend to specialize in tasks and find ways to offload repetitive tasks to automated computer systems. Laboratory scientists, on the other hand, are practical, trained in concrete observations and prefer to accurately describe the specific results of experiments through interpretable analyses.
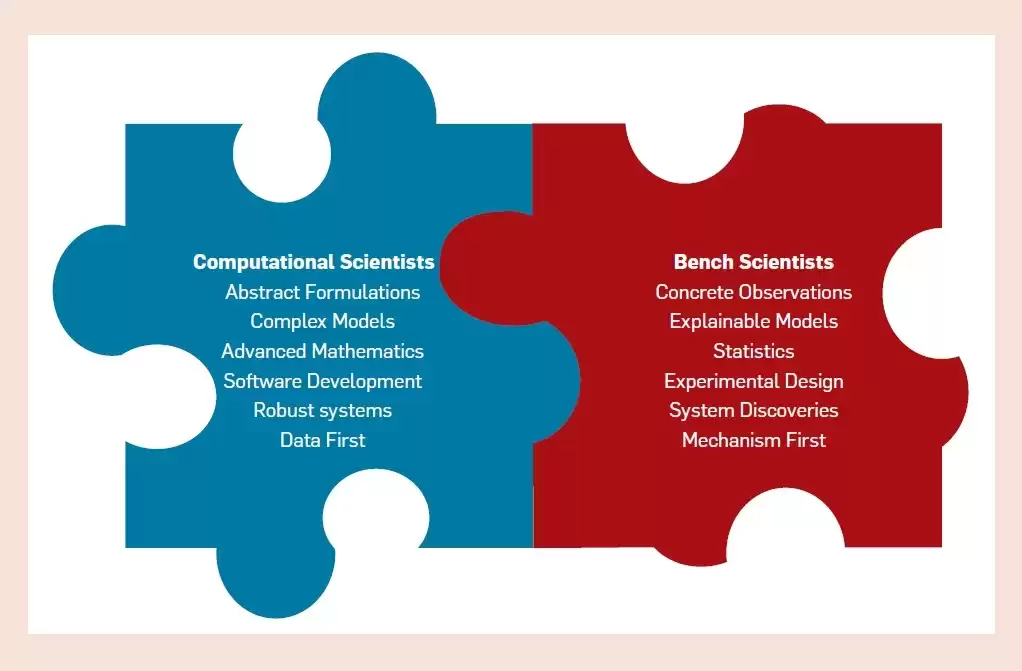
The two worlds have different cultures, which is reflected not only in how the two groups solve problems, but also in what problems they think are worth solving.
For example, there has been a tension between efforts dedicated to building infrastructure that supports general research and efforts dedicated to investigating specific research questions. Computational scientists tend to provide reliable infrastructure that can be used for a variety of projects, while experimental scientists tend to focus on the end goal. Computational scientists prefer to develop mathematical models to explain and predict the behavior of biological systems, while laboratory scientists prefer to generate qualitative hypotheses and test these hypotheses through experiments as quickly as possible (at least when studying microorganisms, since these experiments can be performed quickly in 3-5 days Almost done).
Furthermore, computer scientists are often only excited about lofty goals, such as bioengineering organisms for Mars, writing compilers for life that can create DNA to meet required specifications, reconstructing trees to take desired shapes, and bioengineering dragons in reality. In life, artificial intelligence may be used to replace scientists. Scientists at the lab dismiss such goals as pure “hype” because in previous cases, the computing genre promised a lot but didn’t deliver, preferring to consider only what can be achieved using the current state of the technology.
Solve society’s challenges. The solution to these sociological problems is to encourage interdisciplinary teams and needs. Although we cannot deny that achieving this kind of inclusive environment may be easier in a company (where the team prospers and the team loses) than in an academic environment, where a graduate student or postdoc is often the first author of several papers. Just claim success without the need for integration with other disciplines.
One possible way to achieve this integration is to create cross-training courses where laboratory scientists are trained in programming and machine learning and computational scientists are trained in experiments. This will bring some valuable, unique and necessary cultural exchanges to both communities. The sooner everyone discovers this, the faster synthetic biology can develop.
In the long term, we need university courses that combine the teaching of biology and bioengineering with automation and mathematics. Although some schools are currently offering such courses, they are only a drop in the bucket so far.
◆Perspectives and opportunities
Artificial intelligence can radically enhance synthetic biology and make its full impact by adding a third axis to the engineering stage space, such as physics, chemistry, or biology. Most obviously, AI can produce accurate predictions in bioengineering results, enabling efficient reverse engineering.
In addition, AI can support scientists in designing experiments and choosing when and where to take samples, a problem that currently requires trained experts. AI can also support automated searches, high-throughput analysis, and hypothesis generation based on big data sources, including historical experimental data, online databases, ontologies, and other technical materials.
Artificial intelligence can increase the knowledge of experts in the field of synthetic biology by allowing them to explore large design spaces more quickly and come up with some interesting “outside the box” hypotheses. Synthetic biology poses some unique challenges for current artificial intelligence solutions. If these challenges are solved, it will enable fundamental advancements in the fields of synthetic biology and artificial intelligence. Designing biological systems essentially relies on the ability to control the system, which is the ultimate test of understanding the basic laws of the system. Therefore, AI solutions that enable synthetic biology research must be able to describe the mechanisms that lead to the best predictions.
Although recent artificial intelligence technologies based on deep learning architectures have changed the way we think about feature engineering and pattern discovery, they are still in their infancy in terms of the ability to reason about and explain their learning mechanisms.
Therefore, AI solutions that combine causal reasoning, explainability, robustness, and uncertainty estimation needs have huge potential impact in this interdisciplinary field. The complexity of biological systems makes it impossible for artificial intelligence solutions based purely on brute force correlation discovery to effectively describe the intrinsic characteristics of the system. A new class of algorithms that smoothly combine physical and mechanical models with data-driven models is an exciting new research direction. We are currently seeing some initial positive results in climate science and computational chemistry, and hope that similar progress will be made in the study of biological systems.
As AI provides tools to modify biological systems, synthetic biology can in turn inspire new AI approaches. Biology inspired fundamental elements of artificial intelligence such as neural networks, genetic algorithms, reinforcement learning, computer vision, and swarm robotics. In fact, there are many biological phenomena that can and are worth simulating using digital technology. For example, gene regulation involves an exquisite network of interactions that not only allows cells to sense and respond to their environment, but also keeps cells alive and stable. Maintaining homeostasis (the state of stable internal, physical, and chemical conditions maintained by living systems) involves producing the right cellular components in the right amounts at the right time, sensing internal gradients, and carefully regulating the cell’s exchanges with the environment. Can we understand and exploit this ability to produce truly self-regulating artificial intelligence or robots?
Another example involves emergent properties (i.e., properties displayed by the system but not by its components). For example, an ant colony behaves and reacts as a single organism, not just the sum of its individual ants. Similarly, consciousness (i.e., the perception or awareness of internal or external existence) is a qualitative characteristic derived from a physical basis (such as neurons). Swarm robots that self-organize and collectively build structures already exist. Can we use general theories of emergence to create hybrids of robotic and biological systems? Could we create consciousness from an entirely different physical substrate, such as a transistor? A final possible example involves self-repair and replication: even the simplest examples of life show the ability to self-repair and replicate. Can we understand the dilemma of self-healing and replicating artificial intelligence arising from this phenomenon?
While this type of biomimicry has been considered before, the beauty of “synthetic biology” is that it provides us with the ability to “tinker” with biological systems to test models and underlying principles of biomimicry. For example, we can now tinker with cellular gene regulation at the genome scale, modify it, and test what exactly accounts for its extraordinary resilience and adaptability. Or we could bioengineer ants and test what colony behavior ensues and how that affects ant survival. Or we could alter the cell’s self-repair and self-replication mechanisms and test the effects of long-term evolution on its ability to compete.
Furthermore, in cell modeling we are able to gain a good understanding of the biological mechanisms involved. Even if you understand how a neural network detects the shape of an eye, it’s unlikely you can understand how the brain does the same thing, but synthetic biology research is different. The predictions of the mechanistic model were not perfect but produced qualitatively acceptable results. Combining these mechanistic models with the predictive power of ML can help bridge the gap between the two and provide biological insights into why some ML models are more effective than others at predicting biological behavior. This insight can lead us to investigate new ML architectures and methods.
Artificial intelligence can help synthetic biology, and synthetic biology can in turn help artificial intelligence. The interaction of these two disciplines in a continuous feedback loop will create a future we cannot imagine now, just as Benjamin Franklin could not imagine his interest in electricity. discoveries that will one day make the Internet possible.
