How can fermentation control software dynamically analyse metabolic pathways, realise intelligent control from ‘experience-driven’ to ‘model prediction-driven’, and significantly improve the efficiency and stability of the fermentation process?
Step 1: Data collection – creating a ‘digital picture’ of fermentation
The first step in getting the software to ‘understand’ the fermentation process is to collect a large amount of key data.
These include parameters such as temperature, pH, dissolved oxygen levels, stirring rates, and substrate concentrations during fermentation, as well as metabolic data such as metabolite concentrations (e.g., glucose, lactate, amino acids), enzyme activity, and ATP/NADH levels.
In addition, researchers integrate genomic, transcriptomic, proteomic and metabolomic data to fully profile the metabolic potential of microorganisms. Through sensors and online monitoring equipment, these data can be transferred to the software in real time, providing a solid foundation for subsequent analyses.
Step 2: Metabolic modelling – from data to model
With this data in hand, the next step was to construct a metabolic network model.
Researchers extract the metabolic pathways of target strains from genomic data and transform them into mathematical models using specialised tools (e.g. COBRApy, metaFlux).
With time series data (e.g., substrate consumption rate, product generation rate), the model can infer changes in metabolic fluxes and identify rate-limiting steps and key metabolic nodes.
Depending on the needs, researchers can choose from different types of kinetic models, flux balance analysis models, machine learning models, or even hybrid models to provide accurate prediction and control strategies for the fermentation process.
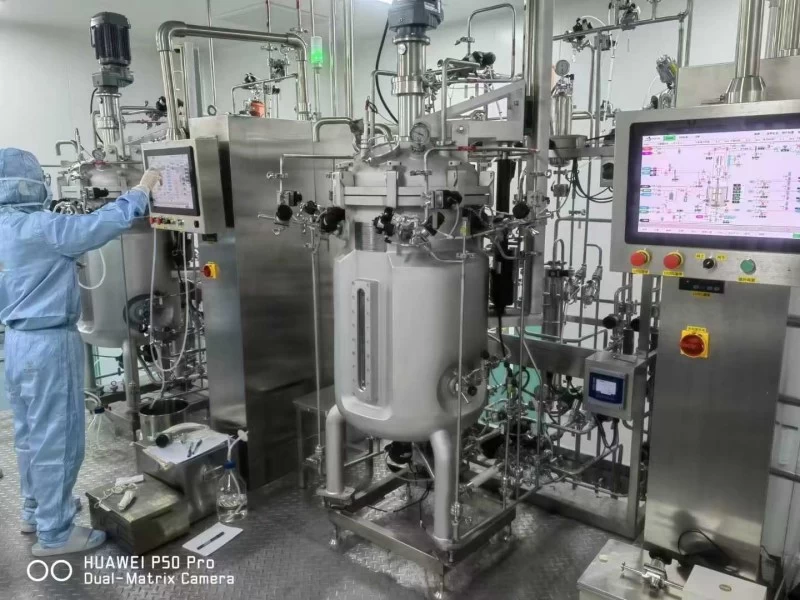

Step 3: Software Integration
Integrating the metabolic model with the fermentation control software is a key step towards intelligence.
Through OPC-UA or API technology, sensor data can be input into the model in real time, and the software dynamically adjusts fermentation parameters according to the model prediction.
At the same time, combined with feed-forward and feedback control strategies, the software can predict and optimise the fermentation process in advance and in real time. In addition, the design of the visualisation interface is also crucial to help operators intuitively understand the fermentation process and the software’s control strategy, thus improving operational efficiency.
Step 4: Model validation and optimisation
The researchers assessed the reliability and accuracy of the model through offline simulations and small pilot fermenter tests.
Based on the experimental data, the model parameters can be further calibrated, secondary metabolic pathways can be simplified, and even reinforcement learning algorithms can be embedded for adaptive optimisation. In the case of yeast production of ethanol, for example, real-time monitoring and modelling predictions have resulted in a 15 per cent increase in ethanol yield and a 10 per cent reduction in the fermentation cycle.
