If you’ve ever interacted with a chatbot like ChatGPT, you’ve experienced the wonders of Large Language Modelling (LLM).
LLM is a machine learning model that is trained with large amounts of natural language data to understand linguistic subtleties, generate new content, and interact with data.
These models can not only come in the form of chatbots, but can also dig deeper into the intrinsic meaning of complex biological datasets.
The Linguistic Fascination of Biodata
Language is viewed as a system of symbols that can be combined in different ways to convey different meanings. Similarly, biological data such as DNA sequences, amino acid sequences, gene expression patterns, etc. can be regarded as a language. They each have a unique ‘vocabulary’ and ‘grammar’ that reflect the various processes and interactions that occur within the organism.
Our bodies use these ‘languages’ fluently and unconsciously, constantly processing instructions and dialogue between cells. Using LLM, researchers are treating this biological data as a language to discover important signals and patterns.
Examples of LLM applications in biology
Genetics and Genomics
The DNA sequence is made up of four basic nucleotides – adenine (A), guanine (G), cytosine (C), and thymine (T) – that form the building blocks of all living organisms. When these sequences are strung together into a complete genome, their variation is linked to complex traits and disease risk factors.
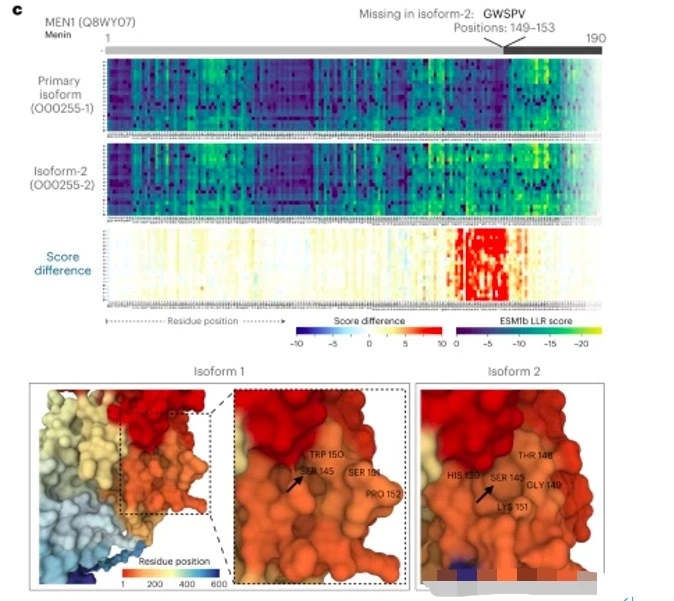
One obvious application of LLM in biology is to study in greater depth how variations in DNA sequences are linked to functional outcomes. For example, Brandes et al. used a protein language model with up to 650 million parameters to predict the phenotypic impact of about 450 million possible error variants in the human genome.
The DNA sequence is made up of four basic nucleotides – adenine (A), guanine (G), cytosine (C), and thymine (T) – that form the building blocks of all living organisms. When these sequences are strung together to form a complete genome, their variation is linked to complex traits and disease risk factors.
One obvious application of LLM in biology is to study in greater depth how variations in DNA sequences are linked to functional outcomes. For example, Brandes et al. used a protein language model with up to 650 million parameters to predict the phenotypic impact of about 450 million possible erroneous variants in the human genome.
These variants are associated with many protein changes, which in turn are linked to disease mechanisms and potential therapeutic targets, and this kind of comprehensive protein-disrupting variant analysis has great potential to enhance human health.
Transcriptomics
Transcriptomic data provide rich insights into understanding the role of RNAs in development and disease and can drive personalised medicine.The application of LLM in analysing single-cell RNA (scRNA) data is b ecoming a powerful tool for understanding biological processes at the cellular level.
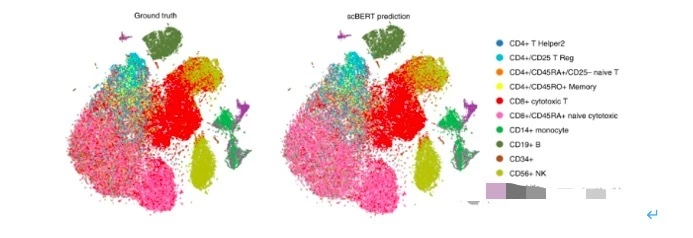
For example, Dr Fan Yang et al. in 2022 successfully developed an LLM-based, scBERT, which is capable of accurately annotating cell types from scRNA sequencing data. dr Theodoris et al. in 2023 developed Geneformer, a scRNA data-based trained transformer model for predicting tissue-specific gene network dynamics in data-constrained situations, accelerating the discovery of key network regulators and candidate therapeutic targets.
Proteomics and Protein Engineering
Understanding the complexity, specific function and drug sensitivity of 3D protein structures is a major challenge, and LLMs have great potential for research in this area.
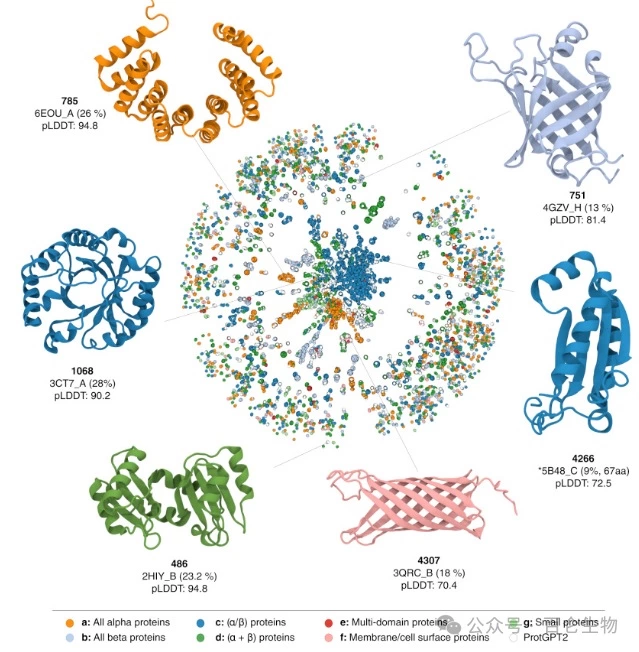
The team led by Dr Madani and Dr Ferruz is developing LLMs called ProGen and ProtGPT2, which are models capable of generating entirely new protein sequences with predictable functions. The output of these models can be further explored for structural significance by tools such as AlphaFold, thus helping to generate biomedically meaningful insights from the rapidly growing protein sequence data.
Small Molecule Drug Discovery and Biochemistry
LLMs not only decode the language of living organisms, they can also play a role in the discovery and optimisation of new drugs. Researchers have transformed libraries of chemical compounds into text-based training datasets to develop chemical language models (CLMs) to predict small-molecule drugs capable of targeting specific proteins in diseases.
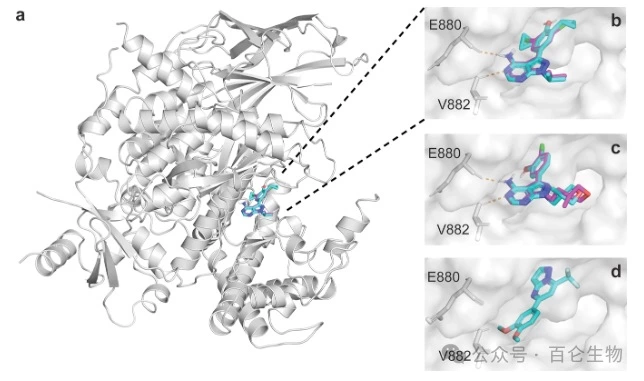
For example, in a 2023 Nature study, Dr Moret et al used CLMs to design a molecule that effectively inhibited abnormalities in the PI3K/Akt pathway associated with many cancer types.
Antibody Evolution and Biologics
The applications of LLMs are not limited to small molecule drug discovery or predicting drug targets; they can also assist in the development of antibodies against diseases. For example, in a Nature Biotechnology 2024 study, Dr Hie et al. used LLMs to guide the laboratory evolution of antibody variants, successfully generating candidates with neutralising activity against Ebola and SARS-CoV-2 viruses.
Overall, large language models have demonstrated their great potential as tools to decipher and reproduce complex relationships and nuances in a wide range of biochemical data forms, from DNA sequences to RNA transcripts to protein sequences and chemical compound libraries.
As LLM becomes increasingly capable of accurately predicting the impact of genetic variants, new therapeutic compounds, and more, scientists will be able to derive actionable insights from data with fewer samples and iterations, rapidly testing more targeted hypotheses. This will ultimately allow researchers to focus on what they do best: asking new questions and imagining new solutions.
