Genome editing is a disruptive biotechnology with broad implications across the life sciences. In the decade since the first groundbreaking CRISPR-Cas9 research, the field has evolved from random, sporadic editing to precise editing techniques. While base editing and particle editing are effective at making small DNA changes, they cannot edit large cargoes.
As the field of genome editing advances, there is an increasing need for efficient and targeted insertion of large pieces of DNA into the genome of living cells.
Gao Caixia’s group from the Institute of Genetics and Developmental Biology of the Chinese Academy of Sciences (CAS) has developed a new genome editing technology that enables efficient and precise directional insertion of large DNA fragments in plants.
The new technique, called Prime editing-mediated re-combination of opportunity targets (PrimeRoot), combines an optimized double ePPE editing protein previously published by the group with a highly efficient tyrosine site-specific recombination Enzyme Cre. It can achieve one-step precise and directional insertion of large DNA fragments in rice and corn with an efficiency as high as 6%, and has been used to successfully insert DNA fragments up to 11.1kb in length.
The results were published in “Nature Biotechnology”. The article is titled “Precise integration of large DNA sequences in plant genomes using PrimeRoot editors”.
CRISPR-associated transposases, a new genome editing tool, have been shown to generate targeted insertions in prokaryotes, but the process is extremely inefficient in eukaryotes. Therefore, despite the rapid expansion of genome editing tools, it is important to be able to generate precise (defined as predictable and specified) large DNA insertions in plants.
Here we describe PrimeRoot (Prime Editing-Mediated Targeted Recombination), a novel genome editing technique that enables precise insertion of large DNA donors in plants without DSB intermediates (Fig. 1a). In addition, PrimeRoot uses many optimized components to increase the overall editing efficiency of plant genomes, while other methods are mostly optimized in mammalian cells.
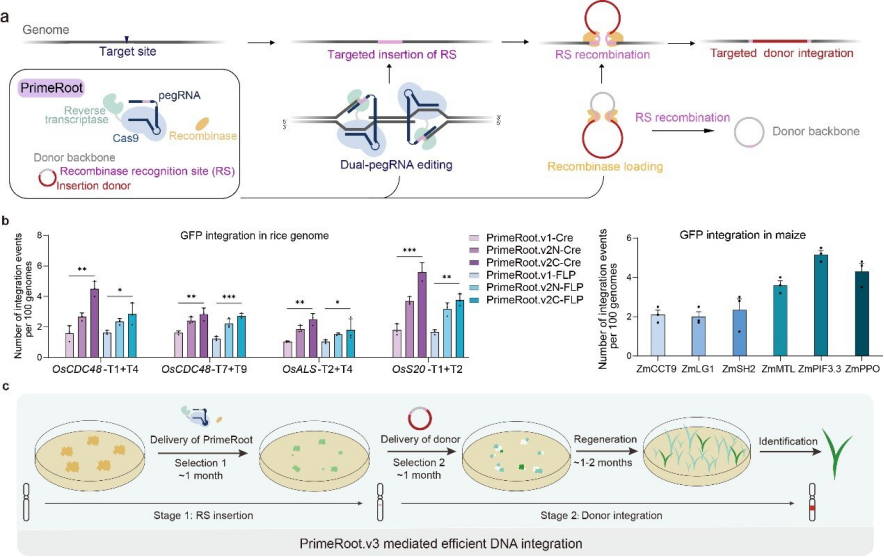
Figure 1: PrimeRoot combines plant-optimized recombinases and enhanced plant PEs to create targeted DNA insertions
Compared with the traditional, imprecise non-homologous end joining strategy, PrimeRoot significantly improves the efficiency of inserting 5kb and above long DNA fragments. It is important that insertion events are completely precise and predictable.
Development of a dual-enhanced plant primer editing system for efficient introduction of recombinase sites in rice
To improve insertion efficiency, the researchers introduced the tevoPreQ1 target, which increases primer editing efficiency to generate dual epegRNAs, and the researchers evaluated combinations of PPE+pegRNA, PPE+PegRNA, ePPE+pegRNA, and ePPE+epegRNA within eight The ability of the origin site to precisely insert the Lox66 and FRT1 RS sequences. The combination of ePPE+epegRNA (hereinafter referred to as “dual ePPE”) showed the highest insertion efficiency (up to 50%) (Fig. 1d).
To expand the targeting range of dual ePPEs, we engineered SpG and SpRY Cas9 variants into ePPEs and assessed their editing efficiency at sites containing NGN-containing PAMs. The researchers confirmed the efficacy of dual ePPE by using this editing method at five loci in rice plants, and found that up to 46% of the regenerated plants had precise RS insertions. These results suggest that the dual ePPE system can efficiently target insertion of RS sequences in plants.
Engineering FRT recombinase sites to improve recombination efficiency
When using PrimeRoot FLP, noticed the presence of short repeats in the FRT1 RS (F1) and wondered if these would reduce insertion frequency. Therefore, the researchers designed three FRT1 mutants (F1m1, F1m2, and F1m3) and two truncated FRT1 (tFRT1) sequence mutants (tF1m2 and tF1m3), each of which was modified based on previously identified key residues. Different point mutations were carried in the RS sequence to see if FLP recombination could be enhanced.
Experiments identified combinations of variants that were more potent than wild-type FRT1, and this was confirmed by examining GFP integration at one site in OsALS using ddPCR (Extended Data Fig. Fig. 3h). Whereas the wild-type combination containing F1+F1 resulted in 1.4% integration, F1m2+F1m3 and tF1m1+F1m3 resulted in 3.0% and 4.0% integration efficiencies, yielding 2.1-fold and 2.8-fold improvements, respectively.
In this study, the researchers also demonstrated two specific applications of PrimeRoot editing. PrimeRoot was used to insert an actin promoter (1.4 kb) upstream of the endogenous OsHPPD gene. The introduction of foreign functional elements is an important genetic breeding method to regulate the expression of endogenous genes. Subsequently, PrimeRoot was used for targeted gene insertion in plants. Traditional methods based on Agrobacterium-mediated and particle bombardment lead to random and imprecise insertion events.
Precise targeting of the actin promoter inserted into the 5′UTR of OsHPPD
Many desirable agronomic traits are quantitative, depending on the up- or down-regulation of certain genes, or on tissue-specific expression.
To see whether PrimeRoot could accurately insert a favorable promoter upstream of the target gene, we used PrimeRoot to knock a strong promoter into the 5′UTR of OsHPPD (Fig. 2c). Sixteen pairs of pegRNAs were designed in the 5′UTR and their RS insertion-editing efficiencies in rice protoplasts were compared. T2+T7 was determined to be the best pegRNA pair with a 30% RS insertion frequency (Fig. 2d).
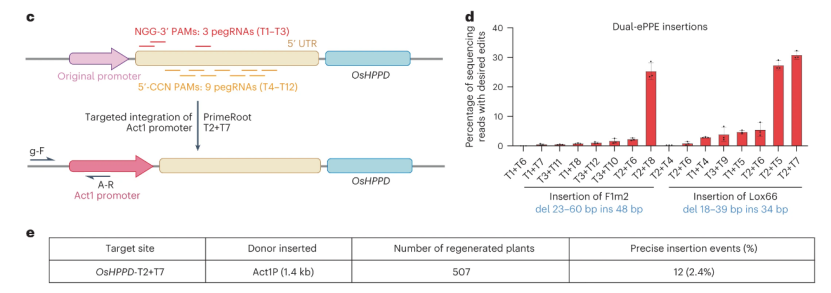
Figure 2: Comparison of targeted DNA insertion mediated by PrimeRoot and NHEJ
Next, the rice actin 1 promoter (Act1P) was inserted into the rice callus by particle bombardment using PrimeRoot and T2+T7. The researchers identified edited plants by amplifying the junction between the genome and the inserted donor sequence, and assessed insertion precision by Sanger sequencing. A total of 12 precise Act1P insertion events (2.4%) were detected in 507 regenerated rice plants (Fig. 2e). These results make PrimeRoot an efficient genome insertion tool for introducing novel genetic regulatory elements into plant genomes for breeding.
To increase the efficiency of PrimeRoot, the researchers also established a sequential transformation system in rice. Compared with a single one-time transformation, this system further improves the editing efficiency by 2 to 4 times, thus achieving an efficiency of up to 8.3% for inserting the actin promoter (1.4kb) and inserting the entire pigmR gene expression cassette (4.9kb) The efficiency is as high as 6.3%.
This technology provides strong technical support for plant molecular breeding and plant synthetic biology research.
